|
|
SME Presentation
Chemotargets
Chemotargets S.L. (www.chemotargets.com) was created in March 2006 as a spin-off initiative from the Chemogenomics Laboratory (http://cgl.imim.es) under the auspices of the Municipal Institute of Medical Research (IMIM-Hospital del Mar: www.imim.es). The company is located within the impressive new premises of the Barcelona Biomedical Research Park (www.prbb.org), in an area that is called to become the "San Diego of the Mediterranean" for its recent growth in the biotechnology sector.

Chemotargets’ core in silico pharmacology platform currently allows for profiling small molecules on almost 1,500 protein targets from the main protein families of therapeutic relevance, including, 837 enzymes, 233 GPCRs, 198 ion channels and transporters, and 32 nuclear receptors. This platform has been internally thoroughly validated and is being exploited through contract service agreements with chemical, biotech, and pharma companies to i) identify new chemical entities with customised target affinity profiles (hit identification), ii) identify potential off-targets at which compounds designed for a particular target may have residual affinity (target fishing), and iii) design chemical libraries directed to particular sets of disease-related targets or entire protein families. The company has also developed proprietary software for the construction of medchem isosteric libraries around known bioactive ligands which, when combined with target profiling, provides a unique means for synthesis planning as well as anticipating all protein targets worth considering in hit and lead optimisation programmes.
In these last three years, Chemotargets has serviced drug discovery projects for 8 pharma companies, 2 chemical companies, and 4 academic institutions and has contributed to the identification of novel hits for known targets, as well as novel targets for known ligands, for many of them. As a result of Chemotargets’ activities, two patents have already been filed and several other projects are currently being pursued internally in those companies to follow up on the hit/target identifications.
For further information on Chemotargets, please visit its website at www.chemotargets.com or send an email to infochemo@chemotargets.com.
By Jordi Mestres
Inte:Ligand

Inte:Ligand is a scientific software and contract research company that was founded 2003 by Prof. Thierry Langer, Dr. Gerhard Wolber and Prof. Hermann Stuppner. Since its foundation, the company has been steadily growing, and now employs 10 scientists, has established several major contract research collaborations and is currently marketing two software products. Inte:Ligand combines expertise in applied molecular modeling supporting medicinal chemistry in form of contract research with software development. In this way, the needs and problems of medicinal chemistry are directly transported to the software developers and can be addressed directly.
Inte:Ligand's key research topic has always been 3D pharmacophore modeling and virtual screening. Their lead product LigandScout, which allows for structure-focused 3D pharmacophore generation has just recently been complemented by ligand-based pharmacophore elucidation [1]. One of the key components of Inte:Ligand's software is a novel, three-dimensional pharmacophore overlay algorithm that takes into account pharmacophoric molecule points and uses pattern recognition for high computational efficiency [2,3]. This pattern recognition bears several advantages over classical multi-point filtering procedures when applied to virtual screening. Another upcoming research area is de-novo design using information from fragment-based screening. The combination of fragments with weak affinity to potent ligands can be performed combining Inte:Ligand's library enumeration technology while geometrically optimizing pharmacophore fitting.
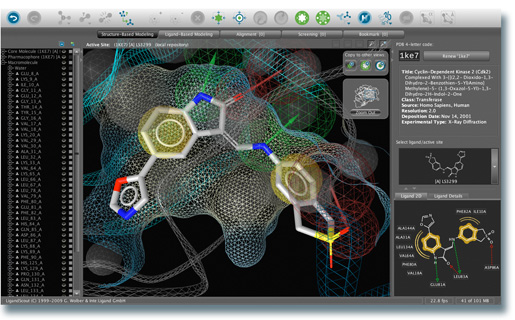
Screenshot of Inte:Ligand's pharmacophore modeling platform LigandScout showing the interactions of a CDK2 protein-ligand complex.
The third research topic of Inte:Ligand is addressing polypharmacology challenges by activity profiling [4]. A collection of approx. 2500 validated 3D pharmacophores have been developed to help in risk assessment in early development stages of drug candidates and are available for parallel virtual screening. They cover 300 unique clinically relevant pharmacological targets originating from major therapeutical classes, such as antiinfectives, cardiovascular, endocrine, gastrointestinal, immunologic, metabolic, neurologic, oncolytic, renal-urologic, and respiratory agents as well as antitargets such as hERG, and members of the cytochrome P450 family.
Inte:Ligand is combining applied research with innovative software development focusing on three-dimensional pharmacophores. Specializing on this area, it has become a key player in the cheminformatics and virtual screening community.
Information and contact:
Dr. Gerhard Wolber
Mariahilferstrasse 74B/11
1070 Vienna, Austria
Web: www.inteligand.com
E-Mail: office@inteligand.com
Selected publications:
- [1] G. Wolber and T. Langer. LigandScout: 3-D Pharmacophores Derived from Protein-Bound Ligands and Their Use as Virtual Screening Filters. J. Chem. Inf. Model; 2005; 45(1); 160-169.
- [2] Efficient overlay of small organic molecules using 3D pharmacophores J. Comput. Aided Mol. Des.; 2007; 20(12); 773-788.
- [3] G. Wolber, T. Seidel, F. Bendix, and T. Langer. Molecule-Pharmacophore superpositioning and pattern matching in computational drug design. Drug Discov Today. (1-2):23-29 (2008).
- [4] T. M. Steindl, D. Schuster, G. Wolber, C. Laggner, T. Langer. High Throughput Structure-based Pharmacophore Modeling as A Basis for Successful Parallel Virtual Screening J. Comput. Aided Mol. Des., 20, 703-715 (2006).
By Gerhard Wolber
|
|

Editor
Gabriele Costantino
Univ. Parma, IT
Editorial Committee
Erden Banoglu
Gazi Univ., TR
Jordi Mestres
IMIM-UPF, ES
Wolfgang Sippl
Univ. Halle-Wittenberg, DE
Kristian Stromgaard
Univ. Copenhagen, DK
Mark Lansdell
Pfizer, UK

Executive Committee
Gerhard Ecker President
Roberto Pellicciari Past-Pres.
Koen Augustyns Secretary
Rasmus P.Clausen Treasurer
Javier Fernandez Member
Mark Bunnage Member
Peter Matyus Member
|







